Lars Kolbowski, Swantje Lenz, Lutz Fischer, Ludwig R. Sinn, Francis J. O’Reilly, Juri Rappsilber
Analytical Chemistry, 2022
10.1021/acs.analchem.1c05266
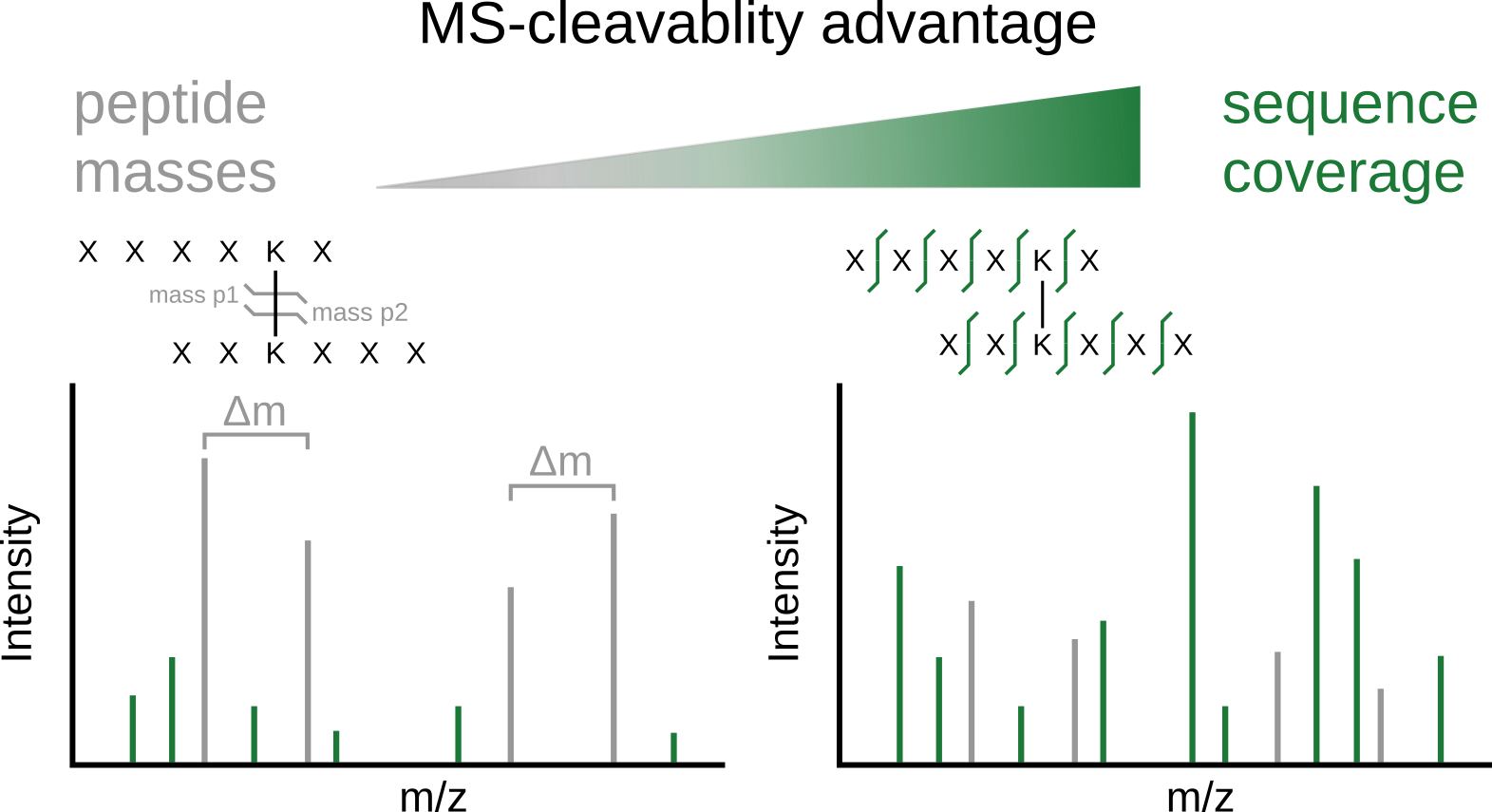
Proteome-wide crosslinking mass spectrometry studies have coincided with the advent of mass spectrometry (MS)-cleavable crosslinkers that can reveal the individual masses of the two crosslinked peptides. However, recently, such studies have also been published with noncleavable crosslinkers, suggesting that MS-cleavability is not essential. We therefore examined in detail the advantages and disadvantages of using the commonly used MS-cleavable crosslinker, disuccinimidyl sulfoxide (DSSO). Indeed, DSSO gave rise to signature peptide fragments with a distinct mass difference (doublet) for nearly all identified crosslinked peptides. Surprisingly, we could show that it was not these peptide masses that proved the main advantage of MS cleavability of the crosslinker, but improved peptide backbone fragmentation which reduces the ambiguity of peptide identifications. This also holds true for another commonly used MS-cleavable crosslinker, DSBU. We show furthermore that the more intricate MS3-based data acquisition approaches lack sensitivity and specificity, causing them to be outperformed by the simpler and faster stepped higher-energy collisional dissociation (HCD) method. This understanding will guide future developments and applications of proteome-wide crosslinking mass spectrometry.