to our understanding how cells work and how diseases can be cured.
and interactions in cells and make them universally accessible.


and interactions in cells and make them universally accessible.


and interactions in cells and make them universally accessible.


and interactions in cells and make them universally accessible.


LATEST PUBLICATIONS
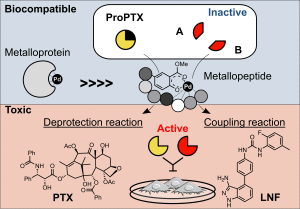
Dual-Bioorthogonal Catalysis by a Palladium Peptide Complex
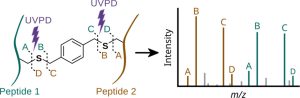
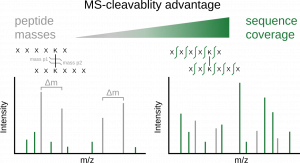
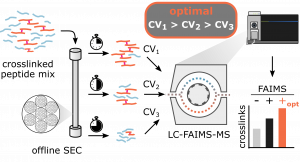
Improved Peptide Backbone Fragmentation Is the Primary Advantage of MS-Cleavable Crosslinkers
LAB NEWS
We’re thrilled to share that we’ve officially begun the My Green Lab Certification process! 📷 This marks an important step toward making our laboratory operations more sustainable and environmentally conscious.
Funding
We gratefully acknowledge support by our funders especially the Wellcome Trust for their willingness to fund us when our vision was daring but the prospects of ever reaching our goals still far remote. We are also deeply indebted to Edinburgh for providing a very supportive research environment and many great colleagues for being the way they are. This allowed us to ultimately overcome the key challenges of our approach. We also thank the many gifted students of Biotechnology at the TU Berlin for their curiosity and interest in technology development and the TU Berlin for providing a space to tune and scale our approach.